Watching the immune system in action has been key to Ziv Shulman’s discoveries about how the body’s peripheral lymph system and gut immune system make antibodies precisely targeted to fight pathogens in very distinct ways. Shulman has invented and applied cutting-edge visualization tools and techniques to show how the evolutionary selection process generates the most protective antibodies against a particular pathogen.
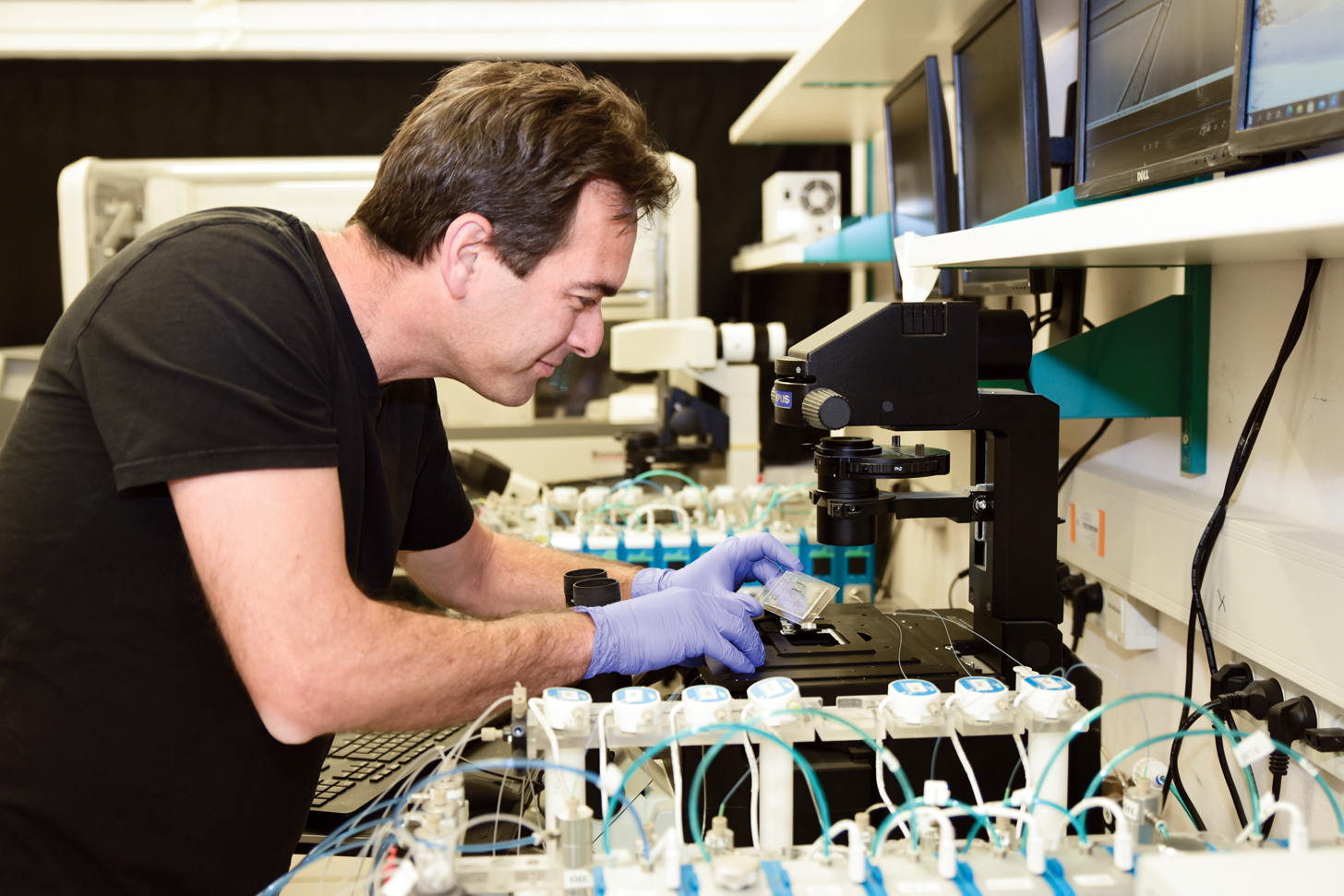
“Imaging through live microscopy is very exciting and stimulating because you get to watch the antibody immune response and dynamics with your own eyes,” says Shulman, principal investigator and head of The Shulman Lab, which focuses on cellular dynamics and molecular regulation of the adaptive immune response.
As an Azrieli Faculty Fellow in the Weizmann Institute’s Department of Immunology from 2016 to 2020, Shulman saw an urgent need for a better way to visualize the poorly understood gut immune system. The specialized immune niches, or sites, within the system’s lymphoid organs are so small and well hidden that scientists have found it difficult to study them with standard imaging methods.
“I thought, if you can image a whole brain in neuroscience, why can’t you image the whole organs of the immune system? We learned how to make the gut lymphoid organs transparent using light-sheet fluorescence microscopy and capture all of them to visualize the gut immune system in a more holistic way at single-cell resolution,” Shulman says. “In one shot, you see everything, and we were able to capture small immune niches that are difficult to detect using conventional microscopy. That was a big advance, which allowed us to ask important questions about how the immune response in the gut works.”
In the breakthrough paper “BCR affinity differentially regulates colonization of the subepithelial dome and infiltration into germinal centers within Peyer’s patches” (Nature Immunology, March 2019), Shulman and his colleagues described using whole-organ imaging to show how the gut immune system’s B cells—a type of white blood cell—play by a different set of rules than those in the peripheral lymph system. They visualized the gut lymphoid organs of mice that had been immunized orally to reveal the gut immune response in action. While the peripheral immune system aims to select and produce the most effective, high-affinity antibodies quickly, the gut system antibody response involves a slower two-stage process.
“In the first stage, affinity-based selection is delayed, and both low- and high-affinity antibodies are produced. For the second stage, the immune cells must migrate into niches in which the proper antigens have accumulated over time, so that affinity training can take place to produce high-affinity antibodies,” Shulman says. “But unlike in the peripheral lymph system, the antigen levels within the gut immune system’s niches are mostly too low to stimulate efficient antibody generation.” He discovered that an area of the gut’s lymphoid organs, known as the subepithelial dome, plays a crucial role in their antibody immune response. These findings could help scientists design new and better oral vaccines.
“Our study suggests that an antigen needs to be targeted to the subepithelial dome to trigger an effective immune response. Good targeting will increase the dose of the vaccine and promote a more effective immune response in the gut. Better targeting can be used in vaccinations against pathogens that enter through mucosal tissues, such as rotavirus, HIV, polio, and bacteria like salmonella,” he says.
Shulman’s interest in immunology research was sparked in 1999 after he persuaded an Israeli biotech company to hire him as a technician. “I got to hang around the lab and see what they were doing. Their goal was to cure spinal cord injuries using the body’s immune system. It was fascinating, and I wanted to be like that,” Shulman says. The experience inspired him to start a bachelor’s degree in animal science at the Hebrew University of Jerusalem in 2001. His subsequent cutting-edge research on immune cell migration and adhesion at the Weizmann Institute earned him prizes from the Feinberg Graduate School for outstanding achievements, once as an MSc student in 2006 and again as a PhD student in 2011.
While completing his PhD research, Shulman used live imaging microscopy to learn about how immune cells pass through cell layers lining blood vessels. Shulman’s and the lab team’s findings were published in the article “Transendothelial migration of lymphocytes mediated by intraendothelial vesicle stores rather than by extracellular chemokine depots” (Nature Immunology, December 2011). “I could visualize the immune cells’ tiny, sticky legs moving like a millipede. Live imaging was key to the discovery, and you can’t see it by using traditional microscopy that produces static pictures,” explains Shulman, who did postdoctoral research at Rockefeller University in New York City to learn more advanced imaging techniques and observe how high-affinity antibodies are formed.
Forming efficient antibodies against specific pathogens involves a biological selection process called antibody affinity maturation, in which B cell antibody genes randomly mutate. During his postdoc, Shulman developed a novel system to visualize and analyze how T cells and B cells interact in real time to form the best antibodies against future pathogens in response to vaccination. He used intravital two-photon laser scanning microscopy and automated quantification algorithms to shed light on the process.