When it comes to artificial intelligence, the buzz is all about large language models (LLMs), those massive, energy-hungry algorithms that, today, can write a middling undergraduate essay and, someday, will develop full sentience — at least according to what tech CEOs keep telling us.
But are they overhyped? Every day we encounter new reports about the disruptive — and unproductive — roles that LLMs are playing in our lives. We read about the AI lawyer that invents case law, or the AI book reviewer that recommends nonexistent books. We don’t yet know what LLMs are reliably good for, but we do know one thing: they’re hogging the media attention.
Meanwhile, the revolution may be quietly happening elsewhere. In scientific disciplines ranging from materials discovery to neurology to genetics, machine-learning technologies — usually algorithms that manage large, complicated datasets — have become essential partners in labs around the world. (Machine-learning technologies are a subfield of artificial intelligence.)
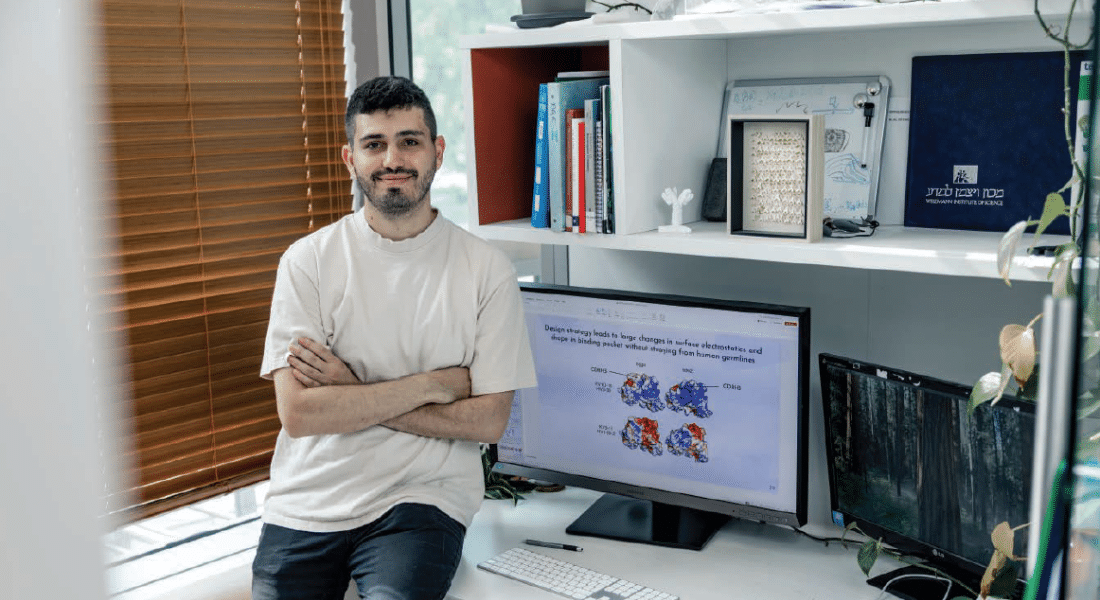
The superpower of machine learning algorithms is their ability to spot correlations that human brains cannot. “It’s difficult for people to pick out complicated non-linear relationships,” says Ariel Tennenhouse, an Azrieli Graduate Studies Fellow and PhD candidate in molecular biology at the Weizmann Institute of Science, in Rehovot, Israel. “If I’m looking at a graph, there has to be an obvious visual trend there or I’ll probably miss it. And I can only handle graphs with two — or, at most three — axes. Machine learning, however, can understand things on a multidimensional plane.”